 
The proportional-odds and multinomial-logistic regression models differ in two ways: the po model has an equal-coefficients (parallel-regressions) assumptions (except for intercepts) and so has fewer parameters the po model models cumulative logits, while the mnl. Here is a small example: library(nnet)ĭf <- data. Dear Steve, The short answer is 'no,' but the test you propose is in my experience usually a close approximation to a valid test. I am new to using to R and to this community, so my apologies if this question is very basic or is missing something essential. Lets use the data from the table and create our Scatter plot and linear regression line: Regression. Lastly, I tried using the mnlAveEffPlot function from the DAMisc package, which works, but isn't as customisable as I would like. > methods (confint) 1 fault confint.glm confint.lm confint.multinom 5 s. ci <- confint (test, level0.95) Note that confint is a generic function and a specific version is run for multinom, as you can see by running. I have also tried MNLpred package and its mnl_pred_ova function as an alternative, but my predictors are in log scale, and it doesn't seem to be able to deal with these - I get the following error: Error in eval(parse(text = paste0("data$", xvari))) : Simply use the confint function on your model object. However, (and I'm sure this is my error somewhere) when I run this code I get the following error:Įrror in FUN(X]. (I also tried the MNLpred package and its mnl_pred_ova function as an alternative, but this one doesn't seem to be able to deal with factors.) I couldn't find any example for the use of ggeffects with multinom, so I'd be grateful for any suggestion that works for numerical and factor predictor variables. If I plot the same data with effects(), I do get the CIs. I have a dataset which consists of Pathology scores (Absent, Mild, Severe) as outcome variable, and two main effects: Age (two factors: twenty / thirty days) and Treatment Group (four factors: infected without ATB infected + ATB1 infected + ATB2 infected + ATB3). 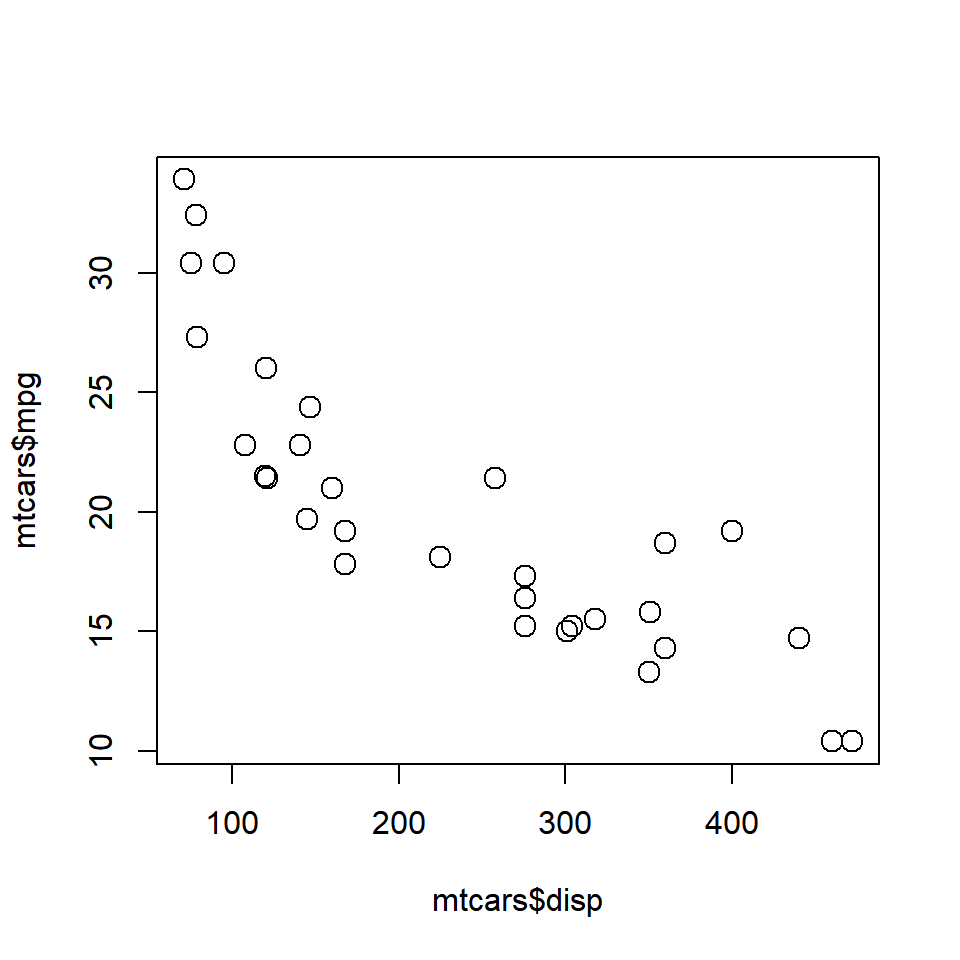
Though ggeffects() should be compatible with multinom, the plot does not display confidence intervals. How do I get p-values using the multinom function of nnet package in R. I have a question regarding the ggeffects package and its use with multinom functions (from nnet package): I am trying to plot marginal effects for a multinomial regression model. If you aren’t using a tool like vtreat in your data science projects: you are really missing out (and making more work for yourself).Plotting confidence intervals with ggeffects for multinom Machine Learning and Modeling.vtreat helps with project blocking issues commonly seen in real world data: missing values, re-coding categorical variables, and dealing high cardinality categorical variables.To use vtreat you only have to work through one introduction (the one helping with the task you have at hand in the language you are using). One way to visualize the results of a multinomial model is simply to plot our fitted values for yon top of our original data: plot(y x, col rgb(0, 0, 0, 0.05), pch 19) lines(newdatax, p1, col rgb(1, 0, 0, 0.75), lwd 5) This plot shows that as we increase along x, observations are first likely to be in y2, then 圓, and finally y1. 
That is now 8 introductions to start with. Multinomial classification: R multinomial classification example, Python multinomial classification example.Unsupervised data preparation: R unsupervised example, Python unsupervised example.Classification: R classification example, Python classification example.Regression: R regression example, Python regression example. 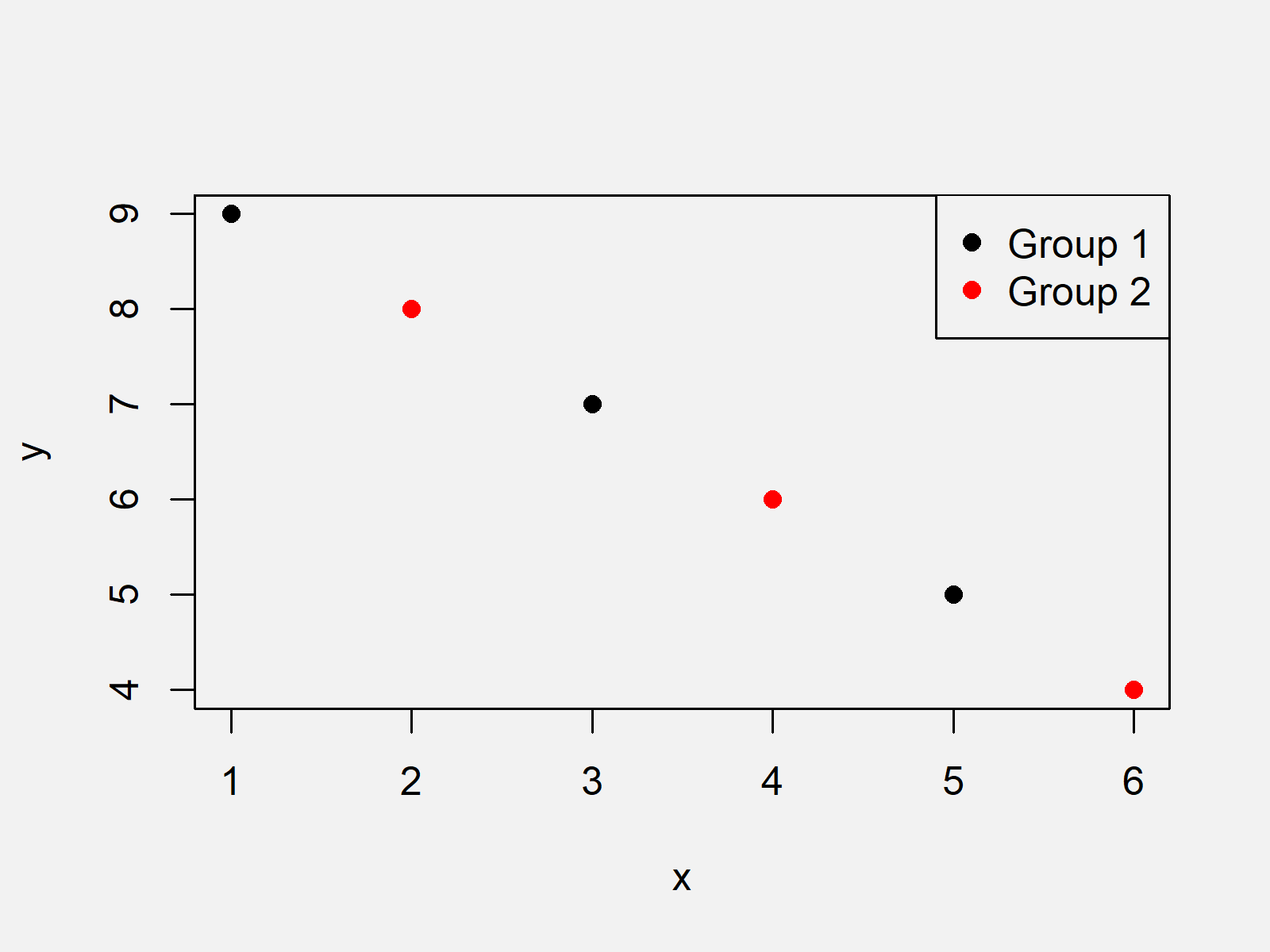
So we now have good introductions on how to use vtreat to prepare data for the common tasks of: R - R code to easily make regression coefficient plots from JAGS/BUGS. And I have finally finished porting the documentation to R vtreat. R - fit a Bayesian multinomial logit model, using Austrian voting data in R using. You can specify the behaviour - na.omit and na.exclude: returns the object with observations removed if they contain any missing values differences between omitting and excluding NAs can be seen in some prediction and residual functions - na.pass: returns the object unchanged - na. Nina Zumel finished some great new documentation showing how to use Python vtreat to prepare data for multinomial classification mode. New vtreat Documentation (Starting with Multinomial Classification) Estimate of the Posterior Distribution of the Price Coefficient (Right. Plotting predicted probabilities and confidence intervals using ggplot2. Figure 2: Time-series Plot of Three Independent Markov Chains (Left Panel) and A Density. Data science advising, consulting, and training 81), also available in the R package arm. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
